Abstract
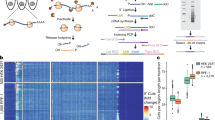
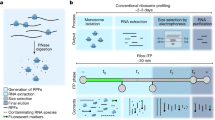
Cellular diversity and architectural complexity create barriers to understanding the function of the mammalian CNS at a molecular level. To address this problem, we have recently developed a methodology that provides the ability to profile the entire translated mRNA complement of any genetically defined cell population. This methodology, which we termed translating ribosome affinity purification, or TRAP, combines cell type–specific transgene expression with affinity purification of translating ribosomes. TRAP can be used to study the cell type–specific mRNA profiles of any genetically defined cell type, and it has been used in organisms ranging from Drosophila melanogaster to mice and human cultured cells. Unlike other methodologies that rely on microdissection, cell panning or cell sorting, the TRAP methodology bypasses the need for tissue fixation or single-cell suspensions (and the potential artifacts that these treatments introduce) and reports on mRNAs in the entire cell body. This protocol provides a step-by-step guide to implement the TRAP methodology, which takes 2 d to complete once all materials are in hand.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Heiman, M. et al. A translational profiling approach for the molecular characterization of CNS cell types. Cell 135, 738–748 (2008).
Mellen, M., Ayata, P., Dewell, S., Kriaucionis, S. & Heintz, N. MeCP2 binds to 5hmC enriched within active genes and accessible chromatin in the nervous system. Cell 151, 1417–1430 (2012).
St. John, P.A., Kell, W.M., Mazzetta, J.S., Lange, G.D. & Barker, J.L. Analysis and isolation of embryonic mammalian neurons by fluorescence-activated cell sorting. J. Neurosci. 6, 1492–1512 (1986).
Wang, S., Roy, N.S., Benraiss, A. & Goldman, S.A. Promoter-based isolation and fluorescence-activated sorting of mitotic neuronal progenitor cells from the adult mammalian ependymal/subependymal zone. Dev. Neurosci. 22, 167–176 (2000).
Tomomura, M., Rice, D.S., Morgan, J.I. & Yuzaki, M. Purification of Purkinje cells by fluorescence-activated cell sorting from transgenic mice that express green fluorescent protein. Eur. J. Neurosci. 14, 57–63 (2001).
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Lobo, M.K., Karsten, S.L., Gray, M., Geschwind, D.H. & Yang, X.W. FACS-array profiling of striatal projection neuron subtypes in juvenile and adult mouse brains. Nat. Neurosci. 9, 443–452 (2006).
Marsh, E.D., Minarcik, J., Campbell, K., Brooks-Kayal, A.R. & Golden, J.A. FACS-array gene expression analysis during early development of mouse telencephalic interneurons. Dev. Neurobiol. 68, 434–445 (2008).
Cahoy, J.D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008).
Sugino, K. et al. Molecular taxonomy of major neuronal classes in the adult mouse forebrain. Nat. Neurosci. 9, 99–107 (2006).
Hempel, C.M., Sugino, K. & Nelson, S.B. A manual method for the purification of fluorescently labeled neurons from the mammalian brain. Nat. Protoc. 2, 2924–2929 (2007).
Okaty, B.W., Miller, M.N., Sugino, K., Hempel, C.M. & Nelson, S.B. Transcriptional and electrophysiological maturation of neocortical fast-spiking GABAergic interneurons. J. Neurosci. 29, 7040–7052 (2009).
Barres, B.A., Silverstein, B.E., Corey, D.P. & Chun, L.L. Immunological, morphological, and electrophysiological variation among retinal ganglion cells purified by panning. Neuron 1, 791–803 (1988).
Luo, L. et al. Gene expression profiles of laser-captured adjacent neuronal subtypes. Nat. Med. 5, 117–122 (1999).
Yao, F. et al. Microarray analysis of fluoro-gold labeled rat dopamine neurons harvested by laser capture microdissection. J. Neurosci. Methods 143, 95–106 (2005).
Surmeier, D.J., Song, W.J. & Yan, Z. Coordinated expression of dopamine receptors in neostriatal medium spiny neurons. J. Neurosci. 16, 6579–6591 (1996).
Toledo-Rodriguez, M. et al. Correlation maps allow neuronal electrical properties to be predicted from single-cell gene expression profiles in rat neocortex. Cereb. Cortex 14, 1310–1327 (2004).
Doyle, J.P. et al. Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135, 749–762 (2008).
Tryon, R.C., Pisat, N., Johnson, S.L. & Dougherty, J.D. Development of translating ribosome affinity purification for zebrafish. Genesis 51, 187–192 (2013).
Stanley, S. et al. Profiling of glucose-sensing neurons reveals that GHRH neurons are activated by hypoglycemia. Cell Metab. 18, 596–607 (2013).
Heiman, M. et al. Molecular adaptations of striatal spiny projection neurons during levodopa-induced dyskinesia. Proc. Natl. Acad. Sci. USA 111, 4578–4583 (2014).
Hupe, M., Li, M.X., Gertow Gillner, K., Adams, R.H. & Stenman, J.M. Evaluation of TRAP-sequencing technology with a versatile conditional mouse model. Nucleic Acids Res. 42, e14 (2014).
Zhou, P. et al. Interrogating translational efficiency and lineage-specific transcriptomes using ribosome affinity purification. Proc. Natl. Acad. Sci. USA 110, 15395–15400 (2013).
Heintz, N. BAC to the future: the use of bac transgenic mice for neuroscience research. Nat. Rev. Neurosci. 2, 861–870 (2001).
Shizuya, H. et al. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc. Natl. Acad. Sci. USA 89, 8794–8797 (1992).
Shizuya, H. & Kouros-Mehr, H. The development and applications of the bacterial artificial chromosome cloning system. Keio J. Med. 50, 26–30 (2001).
Gong, S., Kus, L. & Heintz, N. Rapid bacterial artificial chromosome modification for large-scale mouse transgenesis. Nat. Protoc. 5, 1678–1696 (2010).
Gong, S. et al. Targeting Cre recombinase to specific neuron populations with bacterial artificial chromosome constructs. J. Neurosci. 27, 9817–9823 (2007).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Thomas, A. et al. A versatile method for cell-specific profiling of translated mRNAs in Drosophila. PloS ONE 7, e40276 (2012).
Mustroph, A., Juntawong, P. & Bailey-Serres, J. Isolation of plant polysomal mRNA by differential centrifugation and ribosome immunopurification methods. Methods Mol. Biol. 553, 109–126 (2009).
Watson, F.L. et al. Cell type-specific translational profiling in the Xenopus laevis retina. Dev. Dyn. 241, 1960–1972 (2012).
Zomzely, C.E., Roberts, S., Gruber, C.P. & Brown, D.M. Cerebral protein synthesis. II. Instability of cerebral messenger ribonucleic acid-ribosome complexes. J. Biol. Chem. 243, 5396–5409 (1968).
Neuwelt, E.A., Boguski, M.S., Frank, J.J., Procter-Appich, K. & Levy, C.C. Possible sites of origin of human plasma ribonucleases as evidenced by isolation and partial characterization of ribonucleases from several human tissues. Cancer Res. 38, 88–93 (1978).
Gauthier, D. & Ven Murthy, M.R. Efficacy of RNase inhibitors during brain polysome isolation. Neurochem. Res. 12, 335–339 (1987).
Jung, R., Lubcke, C., Wagener, C. & Neumaier, M. Reversal of RT-PCR inhibition observed in heparinized clinical specimens. BioTechniques 23, 24, 26, 28 (1997).
McQuillen, K., Roberts, R.B. & Britten, R.J. Synthesis of Nascent Protein by Ribosomes in Escherichia coli. Proc. Natl. Acad. Sci. USA 45, 1437–1447 (1959).
Kilkenny, C., Browne, W.J., Cuthill, I.C., Emerson, M. & Altman, D.G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. J. Pharmacol. Pharmacother. 1, 94–99 (2010).
Acknowledgements
We acknowledge members of the Greengard, Heintz and R. Darnell laboratories for their helpful discussions, advice and feedback. This work was supported by grants from The JPB Foundation, The Simons Foundation and National Institute of Mental Health (NIMH) award MH090963 to P.G., and by The Simons Foundation, The Howard Hughes Medical Institute and NIMH award MH090963 to N.H.
Author information
Authors and Affiliations
Contributions
M.H., R.K. and R.J.F. assembled the step-by-step protocol; M.H., N.H. and P.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Heiman, M., Kulicke, R., Fenster, R. et al. Cell type–specific mRNA purification by translating ribosome affinity purification (TRAP). Nat Protoc 9, 1282–1291 (2014). https://doi.org/10.1038/nprot.2014.085
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.085
This article is cited by
-
Ribosome profiling: a powerful tool in oncological research
Biomarker Research (2024)
-
Cell cycle specific, differentially tagged ribosomal proteins to measure phase specific transcriptomes from asynchronously cycling cells
Scientific Reports (2024)
-
Regulation of the hippocampal translatome by Apoer2-ICD release
Molecular Neurodegeneration (2023)
-
Astrocyte-oligodendrocyte interaction regulates central nervous system regeneration
Nature Communications (2023)
-
Activation of TrkB in Parvalbumin interneurons is required for the promotion of reversal learning in spatial and fear memory by antidepressants
Neuropsychopharmacology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.